Article
“Collaborative Patient Portals: Patient Understanding of Numeric Health Information”
Renato Ferreira Leitão Azevedo; educational psychology; Cognition, Lifespan Engagement, Aging, and Resilience Group
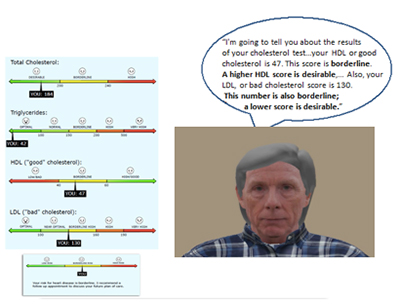
In this project we focus on improving understanding and use of information provided through patient portals to electronic health record (EHR) systems by older adults with diverse numeracy and literacy abilities. Portals are intended to support patient-centered care but are underutilized by older adults because they serve more as information repositories than as collaborative tools for patient education. Self-managing health often hinges on comprehension of numeric information, which challenges lower numeracy patients. Older adults, the most frequent consumers of health information, also have lower numeracy. Patients often interpret numeric information in terms of their goals and knowledge in order to create gist representations organized in terms of evaluative/affective categories that capture the “bottom-line” for their health. Clinicians traditionally help patients create gist by using verbal and nonverbal strategies that contextualize the information. However, clinicians have less and less time to provide this support. While patient portals have the potential to support patient comprehension by providing access to information when needed, portals may not be effective because they eliminate context-supporting gist comprehension. We will help older adults use numeric information by emulating in portal environments best practices from face-to-face communication. Leveraging progress in developing computer-based agents (CA) for human-computer systems, we are developing a CA to serve as an expert clinical intermediary that provides succinct commentary about portal-based test results reinforced by nonverbal cues (facial expressions, tone of voice) to help patients create gist representations that support understanding, engender trust, and encourage them to accomplish self-care goals.
“Molecular Engineering of Sequence-defined Peptide-synthetic Materials for Organic Electronics Using a Molecule Maker”
Eddie Jira, chemical engineering; Beckman Institute Graduate Fellow, Computational Molecular Science Group
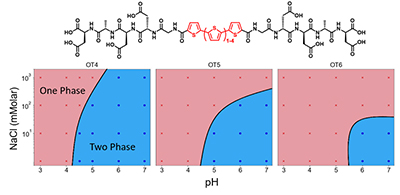
Development of precisely defined conductive polymers for next-generation organic electronic devices is an exciting new direction that promises to unlock high levels of efficiency and performance across a wide range of length scales. This work focuses on development of supramolecular electronic materials, which are composed of precisely defined molecules that can self-assemble into electronically conductive functional hierarchical structures. A major challenge in this field lies in achieving a clear understanding of how primary chemical sequence affects the structural and electronic properties of these devices. From a broad perspective, progress in this area has been hindered by a general inability to readily synthesize large libraries of sequence-defined organic oligomers for this purpose. These challenges are directly addressed by studying the structural and electronic properties of sequence-defined peptide-synthetic biohybrid materials that can be programmed to self-assemble into supramolecular wires. In particular, assembly properties and phase diagrams are determined for a broad class of materials containing oligothiophene cores flanked by sequence-controlled oligopeptides. Moving forward, an automated synthesis platform known as a Molecule Maker is being designed and built to access a broad chemical sequence library, which will be ultimately transferred into a shared facility in the Beckman Institute for researchers to use for various synthetic applications. Overall, this work aims to identify and understand the fundamental relationship between chemical sequence, hierarchical self-assembly, and emergent electronic properties of sequence-defined biohybrid materials.
“Bayesian Modeling of Dynamic Functional Connectivity with Dependent Relevance Determination Priors”
Sameer Manchanda; computer science; Intelligence, Learning, and Plasticity Group
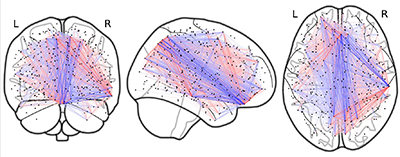
Dynamic functional connectivity (dFC) refers to long-ranging and nonstationary correlations between brain regions observed in functional magnetic resonance imaging (fMRI) studies. Bayesian frameworks allow for model-building that explains data while respecting prior assumptions about how the data was generated. Our research deals with designing and performing inference on Bayesian dFC models. Previous work in our lab has modeled fMRI data using a collection of sparse elementary cognitive units, shared across individuals that vary in activation smoothly over time, with good results on simulated and human connectome project task data. However, the previous approach leaves room for improvement, namely improving modularity and incorporating information from structural priors. I will present ongoing work that encourages the model to learn modular cognitive units, inspired by dependent relevance determination priors, and display and evaluate results on simulated data.
Beckman Institute for Advanced Science and Technology