Article
Characterizing otitis media with optical coherence tomography for improved diagnosis in the primary care and specialist clinic
Guillermo Monroy, Beckman Graduate Fellow, Bioimaging Science and Technology Group
Middle ear infections, commonly referred to as otitis media (OM), affect up to 95 percent of children worldwide. The commonality of this disease poses a significant financial burden on families and healthcare providers to provide time- and cost-effective treatment. Unfortunately, the primary diagnostic instrument for this disease, commonly known as the otoscope, is a relatively simple illuminated magnifier that visualizes the surface of the ear drum. While other diagnostic techniques exist, the results still require subjective interpretation and are often not significantly more accurate than standard otoscopy, at 70 percent for expert physicians.
Our group has miniaturized an optical coherence tomography (OCT) system to fit inside a handheld probe and portable system. For the past several years, observational patient studies have been carried out in collaboration with Carle Foundation Hospital, studying and characterizing OM in roughly 225 patients to date. In this talk, the characterization of OM infection states and diagnosis using OCT will be demonstrated. Past and ongoing projects will be briefly discussed, spanning from the characterization of infection states and their correlation with OCT image features, to machine-learning based classification of OCT data towards the automated diagnosis of OM.
Identifying novel relationships between the developing microbiome and brain using the biomedical piglet model
Austin Mudd, graduate student, Neuroscience Program
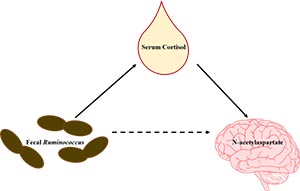
The parallel development of the gut microbiome and the brain early in life provides a window during which highly dynamic changes in the brain could be susceptible to microbial alterations. In fact, mounting evidence suggests dysbiosis of the microbiome may result in commonly observed neurodevelopmental disorders, yet underlying mechanisms remain largely unclear. There is currently a need to characterize variation in microbial composition to identify profiles that may influence specific aspects of brain development. Identifying microbial profiles and products of microbial fermentation that directly affect the etiology of neurodevelopmental disorders may result in improved dietary technologies to combat such disorders. To achieve this goal, an interdisciplinary approach is necessary to understand the complex communication between the gut and brain, which must involve mediating biomarkers. This study integrates multimodal methodology, including magnetic resonance imaging, fecal microbiota measures, and serum biomarkers, to provide novel insight into the complex interactions between the gut microbiota and the brain in early development. Using the biomedical piglet model, we sought to: 1) identify predictive relationships between fecal microbiota and magnetic resonance spectroscopy-derived compounds, 2) identify predictive relationships between fecal microbiota and serum biomarkers, and 3) characterize how serum biomarkers mediate the relationship between fecal microbiota and brain metabolites. Data from our study illuminates a possible connection involved in autism spectrum disorder by providing a novel link between previously established alterations in fecal microbiota, serum biomarkers, and brain metabolites.
Dynamic MR imaging using low-rank tensor models
Bryan Clifford, graduate research assistant, Bioimaging Science and Technology Group
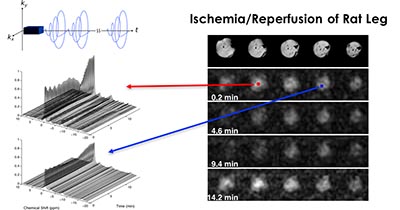
Magnetic resonance (MR) imaging provides a noninvasive window into many aspects of biology. It enables in vivo studies of both structure and function. One of the principle limitations of this technology is its inherently poor trade-off between imaging speed and quality (resolution and noise level), which limits its application to the study of dynamic processes. In this talk I will describe an MR imaging framework which dramatically improves this trade-off through the incorporation of a highly parsimonious low-rank tensor model. I’ll then present some results from the application of this framework to in vivo dynamic oxygen imaging and dynamic spectroscopic imaging.
Beckman Institute for Advanced Science and Technology