Article
Scanning Tunneling Microscopy of Atomically Precise Graphene Nanoribbons Transferred onto H:Si(100)
Adrian Radocea, graduate research assistant in the Nanoelectronics and Nanomaterials Group

ABSTRACT:
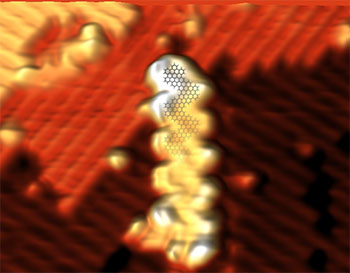
With increasingly demanding miniaturization and performance requirements in the electronics industry, atomically precise graphene materials may play a significant role in high-performance computing. In contrast to conventional top-down manufacturing strategies, the bottom-up approach leads to atomic-level control of the graphene nanoribbon structure enabling fine-tuning of electronic properties.
Graphene nanoribbons are synthesized either on metal surfaces using molecular precursors, or synthesized in solution as demonstrated by the Sinitskii group, our collaborators at the University of Nebraska-Lincoln. Previously, several groups have shown that nanoribbons can be transferred from the metal substrates onto other substrates using wet chemical methods, or solution-synthesized nanoribbons can be drop-cast onto surfaces, but high-quality images and electronic measurements could not be obtained due to problems associated with solvent residue. Our work, recently published in Nano Letters (DOI: 10.1021/acs.nanolett.6b03709), shows that graphene nanoribbons can be cleanly placed onto arbitrary surfaces. Through the use of a dry contact transfer technique developed by the Lyding group, graphene nanoribbons are cleanly transferred under ultra-high vacuum onto silicon, a substrate used for conventional chip manufacturing. Scanning tunneling microscopy (STM) is used to perform nanoscale imaging and electronic characterization of solution-synthesized graphene nanoribbons. Experimental results are verified with computational modeling performed by the Aluru group.
Searching for Image-Based Subtypes of Breast Cancer
Benjamin Chidester, Ph.D. student in Minh Do's Group
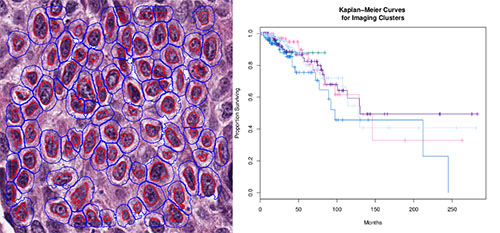
ABSTRACT:
The gold standard for diagnosis and prognosis of breast cancer is analysis of H&E-stained histology slides by a pathologist. We are investigating the existence of meaningful grades or subtypes of breast cancer that can be discovered computationally from H&E histology slides and that can be linked to a patient's outlook or genomic markers. Fundamental to such an investigation are robust and reliable extraction of nuclear and cellular features and accurate segmentation of nuclear and cellular boundaries. To accomplish this, we designed a convolutional neural network to perform accurate nuclear segmentation, improving upon simple thresholding schemes that are often used in such H&E analysis. We also designed a computational pipeline for extracting color, shape, texture, and relational descriptions of nuclei and cells from segmented slides and for visualizing cluster analysis performed on patient image-based feature vectors. I will present results of our investigation on publicly available data from the Cancer Genome Atlas (TCGA), which contains clinical, genomic, and image data for several hundred breast cancer patients.
Multi-Modal Sensor Learning for Continuous Emotion Prediction
Pooya Khorrami, graduate research assistant in the Organizational Intelligence and Computational Social Science Group

ABSTRACT:
Determining a subject’s emotional state automatically from multimedia content is an inherently challenging problem which can be applied to various domains including advertising, medical diagnosis, and human computer interaction. The Audio Video Emotion Challenge (AVEC) 2016 provides a platform for designing and evaluating systems that estimate the arousal and valence states of emotion as a function of time. It also presents the opportunity to investigate multimodal solutions that include audio, video, and physiological sensor signals. Here we will provide an overview of our entry to the AVEC 2016 challenge. Our system consists of high- and low-level features for modeling emotion in the audio, video, and physiological channels. The low-level features model arousal in the audio data with minimal prosodic-based descriptors, while the high-level features are derived from supervised and unsupervised machine learning approaches based on sparse coding and deep neural networks. Predictions generated from each of the input modalities are then combined using a Kalman filter. We not only present results showing how our proposed features outperform the baseline system, but we also demonstrate how using multiple modalities can compensate for instances when one of the input modalities is unreliable or absent.
Beckman Institute for Advanced Science and Technology