Article
This Q&A is part of the Grad Student Visionaries series, a spin-off of our Student Researcher Spotlight that highlights the state-of-the-art equipment housed in Beckman's Microscopy Suite and Visualization Laboratory and the graduate students who use it.
For an animal tasked with racing, jumping, and carrying humans, it’s not uncommon for horses to experience lameness, or a change in gait due to pain or weakness in the limb.
 Sara Moshage
That’s why Sara Moshage, a graduate student in mechanical engineering and graduate fellow at the Beckman Institute, is researching young equine bones; specifically, which
training methods are most successful in strengthening them. Moshage is a researcher in the Tissue Biomechanics Lab working with Mariana Kersh, an associate professor of mechanical science and engineering, and Rohit Bhargava, Founder Professor of Engineering and director of the Cancer Center at Illinois.
Sara Moshage
That’s why Sara Moshage, a graduate student in mechanical engineering and graduate fellow at the Beckman Institute, is researching young equine bones; specifically, which
training methods are most successful in strengthening them. Moshage is a researcher in the Tissue Biomechanics Lab working with Mariana Kersh, an associate professor of mechanical science and engineering, and Rohit Bhargava, Founder Professor of Engineering and director of the Cancer Center at Illinois.
Can you explain your research?
When individuals are young, exercise has lifelong benefits for bone health in both humans and animals. Our goal is to reduce the number of fractures that equine athletes, such as racehorses, experience. We are using computer models of young horse bones to determine which exercises are the most beneficial for strengthening the bone.
What Visualization Lab tool do you use the most, and why?
Amira. Amira is an extremely well-developed tool for analyzing stacks of images (such as CT scan or MRI) and can handle very computationally challenging tasks. Additionally, Amira has a well-written and easy-to-follow manual, along with many free instructional videos on YouTube.
Can you share a technique that’s helpful to you on Amira?
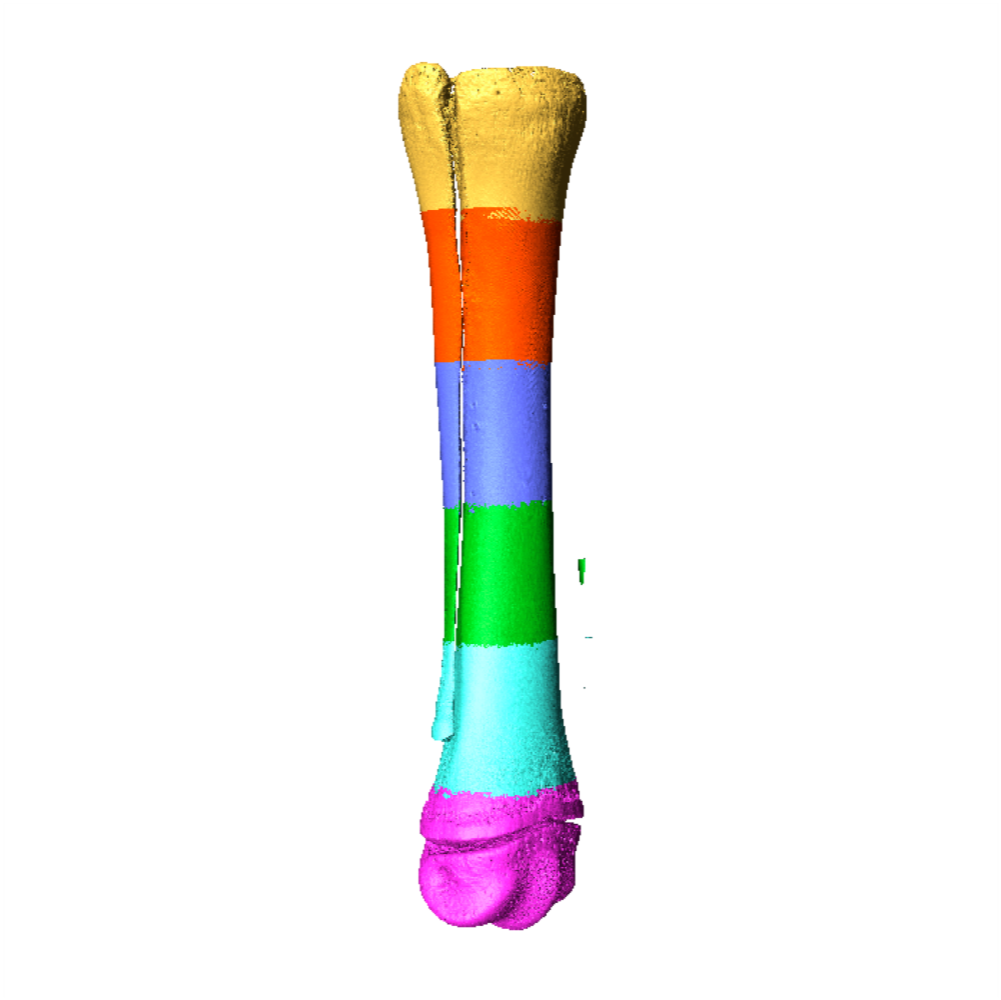 Six individual scans of a juvenile equine cannon bone are merged together to form a "whole bone" scan using Amira. Credit: Sara Moshage.
In the current stage of my project, I'm scanning intact juvenile equine bones in the Rigaku CTLab GX130 microCT scanner. Because of the size of my samples, I have to take several scans to view the whole bone. These scans are positioned to overlap
slightly. Sometimes the bone moves a bit between scans, making it so I can't simply combine the CT images from different scans together and hope it looks like an intact bone. Additionally, I have to remove the 'overlap' between scans.
Six individual scans of a juvenile equine cannon bone are merged together to form a "whole bone" scan using Amira. Credit: Sara Moshage.
In the current stage of my project, I'm scanning intact juvenile equine bones in the Rigaku CTLab GX130 microCT scanner. Because of the size of my samples, I have to take several scans to view the whole bone. These scans are positioned to overlap
slightly. Sometimes the bone moves a bit between scans, making it so I can't simply combine the CT images from different scans together and hope it looks like an intact bone. Additionally, I have to remove the 'overlap' between scans.
This issue of complex image stack alignment in 3D is where Amira really shines. In Amira, it is possible to translate and rotate an image stack around in 3D space in order to align the sections of my intact bone. It is possible to get a decent alignment just by eye, but it's important that the image stacks are aligned as closely as possible so that both the internal structure and the outer contour of the intact bone are smooth and continuous. Amira has a function called Register Images that will sort through two separate image stacks (or CT scans), find the overlapping structure between them, and rotate one of the stacks so that the alignment is perfect. This process can be repeated with fine-tuned parameters in order to have the closest possible alignment between the structure of the two scans. Once this process is repeated for all sections of the intact bone, Amira is able to merge the CT scans into one new image stack that contains the entire bone. The end result is a new image stack of the entire bone where it is all but impossible to see where the image stacks were merged when looking at the bone's internal structure and contours.
Any words of wisdom for future visionaries?
Don’t be afraid to read the manual for the software program you are using. It might be daunting at first, but you’ll always come away from it knowing more than you did before. I’ve learned some really neat skills by reading manuals and looking for instructional videos.
For more information about Beckman Institute's Visualization Laboratory, please visit: https://itg.beckman.illinois.edu/visualization_laboratory/
Sara Moshage uses a software called Amira to merge individual scans of the cannon bone in a juvenile horse into a seamless "whole bone" stack. Video content courtesy of Sara Moshage.
Beckman Institute for Advanced Science and Technology