Article
Researchers at the Beckman Institute for Advanced Science and Technology are working to reconstruct the nervous system of a common crop pathogen, the soybean cyst nematode. Imaging of the nematode is being carried out at the institute’s Microscopy Suite. The work stands out as the first connectome, or map of neural connections, to be assembled for a plant pathogen.
The project is being funded by a grant from the U.S. Department of Agriculture National Institute of Food and Agriculture.
 Nathan Schroeder is imaging cross sections of soybean cyst nematode using scanning electron microscopy.
Nathan Schroeder is imaging cross sections of soybean cyst nematode using scanning electron microscopy.The motivation for this work lies in mitigating crop loss by understanding the environmental signals that lead to soybean cyst nematode infection and intercepting those signals. SCN infects soybean roots by secreting enzymes, such as cellulases, that trick the plant into indefinitely feeding the nematode.
“We know that gland cells in the nematode’s esophagus are responsible for secreting enzymes, but we don’t yet know what the worm senses to cause that secretion or how that sensation is transmitted through the nervous system,” said Nathan Schroeder, an associate professor of crop sciences at the University of Illinois Urbana-Champaign.
Specifically, Schroeder and collaborators aim to map the network of synapses, or nerve cell connections, that surround the SCN esophagus, which serves as the primary feeding structure for the nematode. “Based on the principles of neuroscience, we can make predictions about the strength of connections,” Schroeder said. “One neuron could have multiple synapses with another cell. Once we know those connections, we can determine the most important neurons for functioning of the esophagus and decide which cells to go after to inhibit feeding, which is the ultimate goal of this project.”
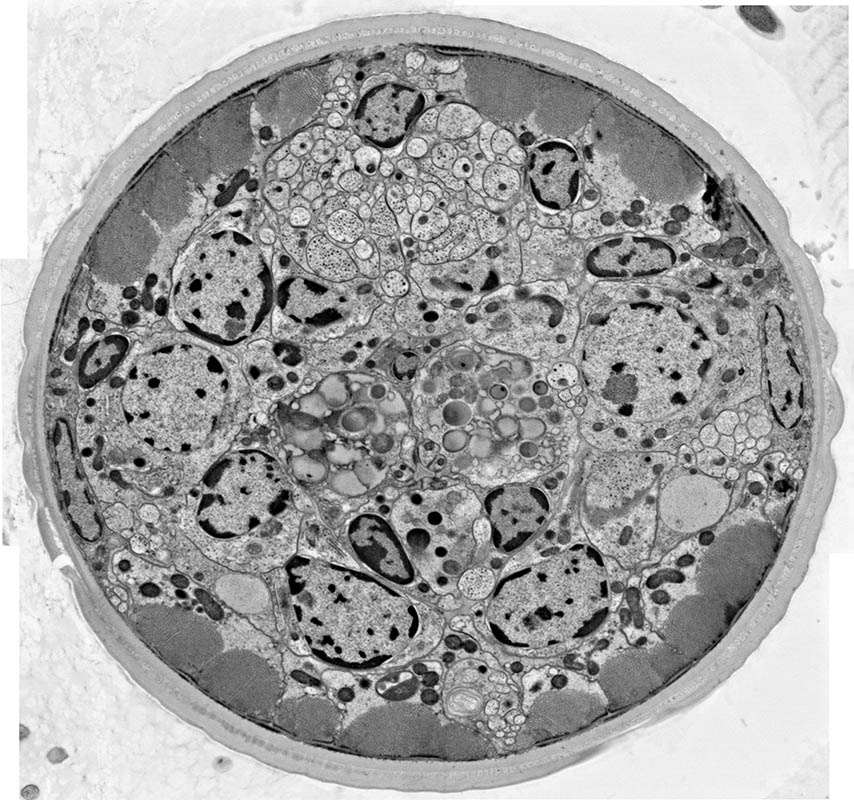 This electron micrograph shows a cross section of soybean cyst nematode. The micrograph was collected using the environmental scanning electron microscope in the Beckman Institute Microscopy Suite.
This electron micrograph shows a cross section of soybean cyst nematode. The micrograph was collected using the environmental scanning electron microscope in the Beckman Institute Microscopy Suite.Very few organisms’ nervous systems have been reconstructed, even partially, since the first connectome was mapped in the 1980s on a nonpathogenic nematode, C. elegans. Reconstructing the C. elegans connectome was a Herculean effort that took over 15 years. “The C. elegans connectome was mapped before digital cameras were invented,” Schroeder said. “Each photographic print had to be hand annotated so the neuronal processes could be followed through over a thousand cross sections. This could only be done for a very simple organism, like C. elegans.”
Fortunately, microscopy advances in the past 30 years allow for a more direct approach to reconstructing the SCN connectome, which when complete will be the first mapped connectome for a parasitic organism. Because individual synapses are too small to be imaged by light microscopy, Schroeder’s group developed a technique for imaging the SCN esophagus using scanning electron microscopy. “Nematodes are already pretty small, but then we use an ultramicrotome to section them into 70 nanometer thick slices,” Schroeder said. “We visualize those thin cross sections with the electron microscope one at a time. One slice represents one of over 2,000 images we would need to reconstruct the nematode’s esophagus. If we image enough slices, we can three-dimensionally map the whole nervous system.”
 Lav Varshney is computationally reconstructing the soybean cyst nematode connectome.
Lav Varshney is computationally reconstructing the soybean cyst nematode connectome.This work is being done in collaboration with Lav Varshney, an associate professor of electrical and computer engineering, who will computationally reconstruct the SCN connectome. “Once the electron micrographs are registered and segmented, and the connectivity pattern between neurons is reconstructed, we are left with a network structure,” Varshney said. “This structure of the neuronal network can then be analyzed using techniques from systems and control theory, to understand how it supports various functions and behaviors.”
The environmental scanning electron microscope Schroeder is using to image SCN is a relatively new addition to the Microscopy Suite, which is managed by Scott Robinson. Robinson received university funding to purchase the ESEM in 2015. Concurrently, Robinson and Schroeder collaborated to acquire funding for a high-pressure freezing and freeze substitution unit, which Schroeder uses to prepare nematode cross sections for electron microscopy. “Both systems were granted by the University, and Nate assigned a graduate student and an excellent electron microscopy technician to work in the Microscopy Suite,” Robinson said. “This collaboration has been very fruitful for each of us.”
Editor’s note:
Although there is not yet a publication associated with this work, relevant prior publications from the Schroeder and Varshney groups can be found in PLOS Pathogens and in PLOS Computation Biology.
The Coordinated Science Laboratory in The Grainger College of Engineering has previously reported on this research. A recent CSL article is online.
The Microscopy Suite is a collection of roughly 30 rooms in the basement of the Beckman Institute. It contains a variety of instrumentation for microscopy and spectroscopy, including fluorescence, atomic force, and electron microscopy.
Beckman Institute for Advanced Science and Technology