Article
Researchers at the University of Illinois Urbana Champaign have developed a new technique that combines label-free imaging with artificial intelligence to visualize unlabeled live cells over a prolonged time. This technique has potential applications in studying cell viability and pathology.
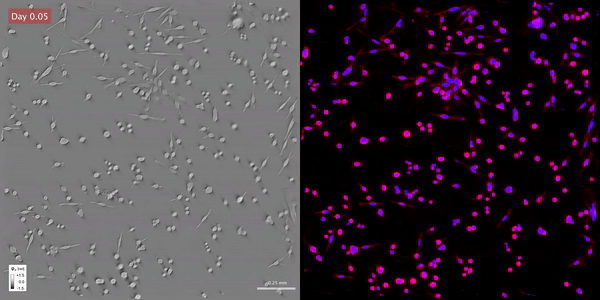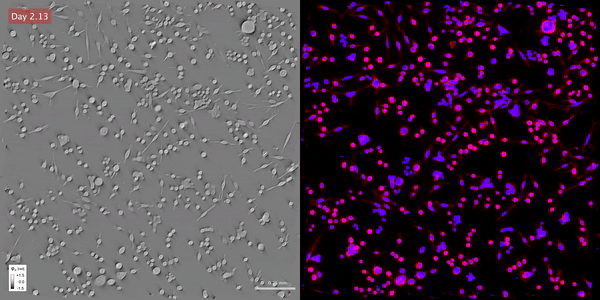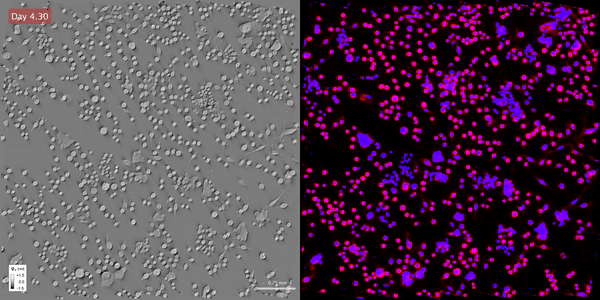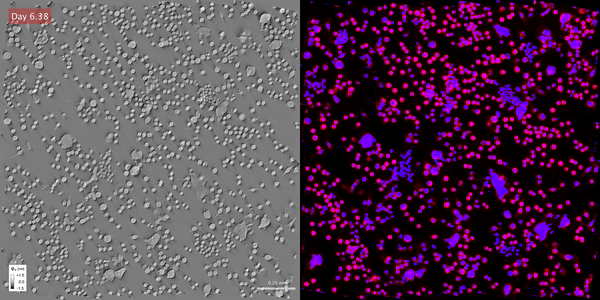 Time-lapse gradient light interference microscopy, or GLIM, left, and phase imaging with computational specificity imaged over seven days.
Time-lapse gradient light interference microscopy, or GLIM, left, and phase imaging with computational specificity imaged over seven days.The study “Phase imaging with computational specificity (PICS) for measuring dry mass changes in sub-cellular compartments” was published in Nature Communications.
“Our lab specializes in label-free imaging, which allows us to visualize cells without using toxic chemicals,” said Gabriel Popescu, a professor of electrical and computer engineering and the director of the Quantitative Light Imaging Laboratory at the Beckman Institute for Advanced Science and Technology. “However, we cannot measure specific attributes of the cell without using toxic fluorescent dyes. We have solved that problem in this study.”
“We had this idea that computational methods could estimate what the sample would look like without actually killing the cells,” said Mikhail Kandel, a graduate student in the Popescu group.
The researchers first imaged the cells over several days using their non-destructive label-free technique. At the end of the experiment, they stained the samples and used deep learning, which is a subset of machine learning, to learn where the fluorescence dyes would be located. “This let us estimate the stain in our initial movies without actually staining the cells,” Kandel said.
“Although AI has been used in the past to create one type of imaging from a different type of staining, we were able to program it to analyze the images in real time,” Popescu said. “Using deep learning, we were able to look at cells that had never been tagged with any dye, and the algorithm was able to precisely locate different parts of the cell.”
“Another advantage of the technique is that we can perform experiments over the span of many days. The cells remain alive even after more than a week,” said Yuchen He, a graduate student in the Popescu group. “This cannot be done with fluorescent dyes since the chemical toxicity might kill the cells.”
“This study highlighted the potential of AI-based techniques to learn complicated models such as the concentration of specific dyes, which goes beyond the capabilities of the naked eye,” Kandel said. “The more we can teach our method to recognize patterns, the more kinds of experiments can be performed without resorting to killing the cells."
The researchers are now trying to adapt deep learning algorithms across different cell lines and biological samples. “Training deep learning models requires a large amount of data because we want to ensure that they work well in different scenarios. Fortunately, our imaging instruments make it easy for us to generate the needed training data in an efficient fashion,” He said.
“These deep learning algorithms can be used for several applications,” Popescu said. “We can assess the cell viability over a long time without labeling the cells, we can differentiate between different cell types in diseases, and we can study different cellular processes.”
The study “Phase imaging with computational specificity (PICS) for measuring dry mass changes in sub-cellular compartments” can be found at https://doi.org/10.1038/s41467-020-20062-x.
Beckman Institute for Advanced Science and Technology