Article
The Beckman Institute Graduate Student Seminar Series presents the work of outstanding graduate students working in Beckman research groups. The seminar starts at noon in Beckman Institute 1005 and is open to the public. Lunch will be served.
Highly-accelerated multi-channel MRI with adaptive and self-calibrating acquisition
Behzad Sharif
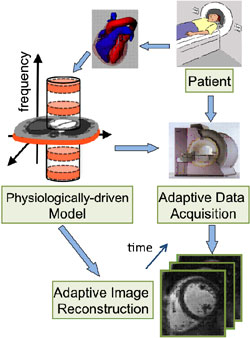
In magnetic resonance imaging (MRI), the computational process of forming an image of the anatomy of interest from the received signals is called “image reconstruction”. This talk focuses on original computational approaches that modify both the data acquisition strategy and the image reconstruction process and achieve more than 10-fold improvement in a combination of performance measures. Thanks to its non-invasive and non-ionizing nature, MRI technology is widely used for anatomical and diagnostic imaging. However, due to MRI’s relatively slow acquisition speed, it typically yields unsatisfactory results in imaging scenarios where high spatial and temporal resolutions are simultaneously desired. We present a method for highly-accelerated MRI of the human heart which overcomes such limitations while enabling “real-time” imaging (i.e., without the need for explicit electro-physiological synchronization). The method incorporates a physiologically-driven model and exploits the redundancy in data acquired by the multiple receiver coils available on standard MRI scanners. Further, an optimized scheme for self-calibration of the image reconstruction process for general multi-channel MRI applications—specifically high resolution brain imaging—is presented. Results of MRI experiments on healthy human volunteers are provided.
Making waves in the stream of consciousness: Eliciting predictable oscillations in visual awareness with rhythmic visual entrainment
Kyle Mathewson
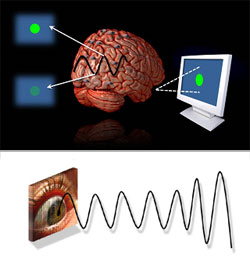
What is the underlying nature of conscious awareness? William James observed introspectively that consciousness, “… does not appear to itself to be chopped into bits…A ‘river’ or a ‘stream’ are the metaphors by which it is most naturally described (James, 1959).” Since this time, however, evidence has accumulated supporting an alternative, discrete nature of perception (Efron 1970; VanRullen & Koch 2003). In a series of studies, we have found evidence that perception fluctuates on a fine temporal scale, as a function of the phase of ongoing neural oscillations. Visual targets presented in the peak of ongoing 10 Hz neural oscillations are visible, while identical stimuli presented in the trough do reach consciousness (Mathewson et al., 2009). Furthermore, it is possible to control these ongoing neural oscillations with rhythmic visual stimulation, eliciting predictably oscillations in consciousness (Mathewson et al., Resubmitted). This thus reveals a method to experimentally control these discrete perceptual snapshots, and provides the first evidence of a causal link between ongoing neural oscillations and the fine grained temporal variations in consciousness.
Template assisted nanofabrication for biological applications
Xindi Yu
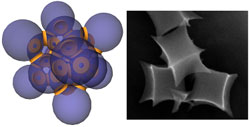
Many biological applications require transportation of bio-materials through the cell membrane. In those applications, nanoparticle based bio-materials provide addition functionality due to their size effect and possible structural complexity. Recent studies have shown that the transport rate of these particles has much to do with their surface curvatures. A challenge to fully understand these effects is to fabricate nanoparticles with the appropriate shape. We developed in our lab a variety of three dimensional template assisted lithography methods that can produce large amount of nanoparticles with well controlled geometry. Collaborating with experts in the bio-material field, we are currently studying the interaction of these particles with cells.
Beckman Institute for Advanced Science and Technology