By
Beckman Institute
Published on
Jan. 21, 2015
Neil Krueger
Tunable Porous Silicon Hybrid Light-Emitting Microcavities
Porous Silicon (PSi) was first discovered in the mid-1950s, but it was a room temperature display of visible luminescence decades later that drew widespread attention. Efforts at all-Silicon (Si) light-emitting devices with PSi ensued. But, despite progress, performance never justified widespread implementation. While interest in PSi for Si-based optoelectronics has subsided, PSi has gained a reputation as a versatile optical material, due largely to its ability to form high-quality optical superlattices through a time-varying current density during electrochemical formation. This enables high-quality factor microcavities that could be coupled with highly efficient emitters and provide a new platform to strongly manipulate emission for applications like lasers and quantum information processing. A hybrid structure composed of a well-defined, high-quality emitting cavity layer between PSi photonic crystal mirrors is particularly attractive to bridge the gap between Si photonics and optoelectronic devices. We have shown that such PSi-based hybrid photonic architectures can be achieved with a modified transfer-printing technique.
Giang-Chau Ngo
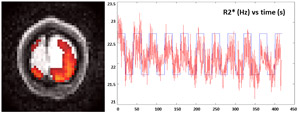
Mapping the Brain and T2*
Magnetic resonance imaging relies on a uniform magnetic field in the brain. When this uniformity is disrupted, there are several effects that can impact an experiment, resulting in image distortions and signal losses. One crucial aspect that is often overlooked is the impact of magnetic field inhomogeneity on the echo time shift of the acquisition. Magnetic field inhomogeneity creates magnetic gradients that interact with imaging gradients used in MRI and have an effect on how the image is acquired. These acquisition distortions can be very adverse to functional MRI (fMRI), since Blood Oxygen Level Depend (BOLD) contrast and the BOLD sensitivity rely on the echo time. However, by taking into account magnetic field gradients, the effect of the echo time shifts can be eliminated. In this work, brain function can be robustly and quantitatively assessed by estimating T2*or R2* directly, a parameter that reflects the brain function. An iterative method has been developed based on a multi-echo spiral sequence, which can accurately quantify R2* despite field inhomogeneity.
Itamar Livnat
A D-amino Acid-containing Neuropeptide Discovery Funnel
Neuropeptides are cell-cell signaling molecules in the nervous system that are derived from enzymatic protein cleavage; the activity of these neuropeptides can be targets for pharmaceutical intervention. These proteins are assembled using exclusively L-amino acids. Thus, it was assumed that D-amino acids, mirror images of L-amino acids, did not naturally occur in animal proteins. However, D-amino acids are found in animal proteins, including in neuropeptides. These D-amino acids are incorporated after assembly of the initial protein, forming a D-amino acid-containing peptide, or DAACP. However, DAACPs are difficult to detect because unlike other peptide modifications, no mass change is associated with them. Thus, I have been developing a methodology for detecting these DAACPs based on their unique properties. Step 1, potential DAACPs can be identified by their resistance to degradation by peptidases. Step 2, I target potential DAACPs from Step 1 for D-amino acid analysis, breaking the peptides into their component amino acids and assaying their amino acid content for D-amino acids. Step 3, DAACPs are confirmed by their bioactivity and matching other properties to synthetic peptide standards that match the suspected sequence of the DAACP. Through this method, non-targeted discovery of this very subtle modification can be carried out.